Background#
This section contains explanations on why the project templates look the way they do. This includes a short explanation of the content of the pre-installed example, the basic design rationale, discussion of the workflow, the directory structure we chose, and a little background on directed acyclic graphs and pytask. There is not much reference to code or a particular programming language here, this is relegated to the next section.
Running example#
The example project that will be installed with the templates is a simple empirical project. Its abstract might read:
This paper estimates the probability of smoking given age, marital status, and level of education. We use the stats4schools Smoking dataset and run a logistic regression. Results are presented in this paper; you may also want to consult the accompanying slides.
We can translate this into tasks our code needs to perform:
Clean the data
Estimate a logistic model
For each of the categorical variables, predict the smoking propensity over the lifetime
Visualize the results
Create tables with the results
Include the results in documents for dissemination
In these templates, we categorize these tasks into four groups:
Data Management: task 1
Analysis: tasks 2 & 3
Final: tasks 4 & 5
Paper: task 6
Naturally, different projects have different needs. E.g., for a simulation study, you might want to discard the data management part. Doing so is trivial by just deleting the respective directory (once you do not need the example any more). For most economics research projects, however, the basic structure has proven to strike a good balance between keeping related code in one place and dividing it up into chunks of manageable size.
The remainder of this section provides much more detail on why we made these choices.
Design Rationale#
The design of the project templates is guided by the following main thoughts:
Separation of logical chunks: A minimal requirement for a project to scale.
Only execute required tasks, automatically: Again required for scalability. It means that the machine needs to know what is meant by a “required task”.
Reuse of code and data instead of copying and pasting: Else you will forget the copy & paste step at some point down the road. At best, this leads to errors; at worst, to misinterpreting the results.
Be as language-agnostic as possible: Make it easy to use the best tool for a particular task and to mix tools in a project.
Separation of inputs and outputs: Required to find your way around in a complex project.
How to Organize the Workflow?#
A naive way to ensure reproducibility is to have a workflow script (do-file, m-file, …) that runs each file one after the other. One way to implement that for the above setup would be to have code for each step of the analysis and a loop over both categorical variables within each step:
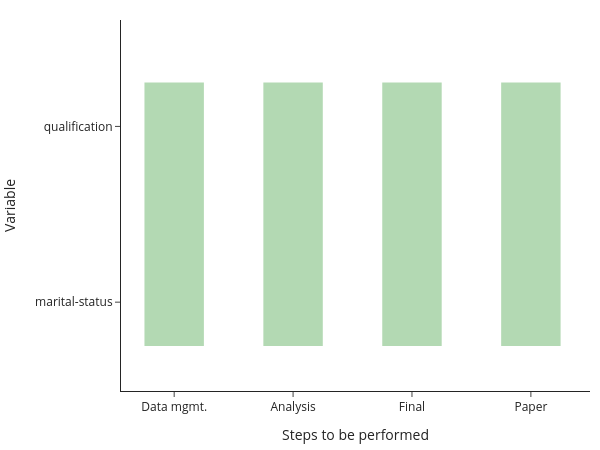
You will still need to manually keep track of whether you need to run a particular step after making changes, though. Or you run everything at once, all the time. Alternatively, you may have code that runs one step after the other for each variable:
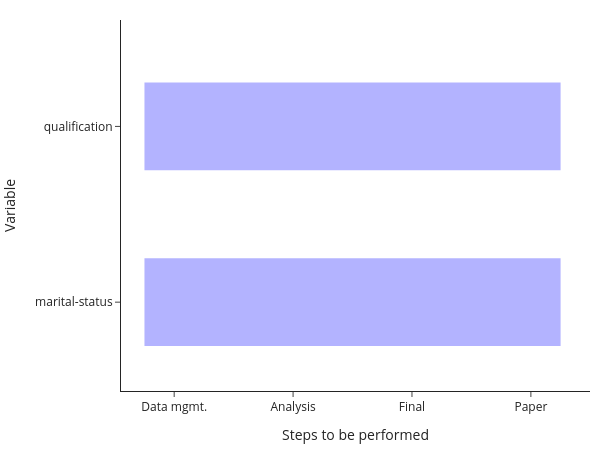
The equivalent comment applies here: Either keep track of which model needs to be run after making changes manually, or run everything at once.
Ideally though, you want to be even more fine-grained than this and only run individual elements. This is particularly true when your entire computations take some time. In this case, running all steps every time via the workflow script simply is not an option. All my research projects ended up running for a long time, no matter how simple they were…
The figure shows you that even in this simple example, there are now quite a few parts to remember:
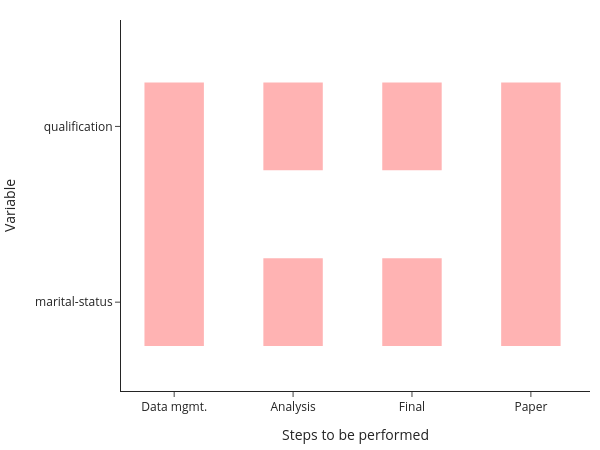
This figure assumes that your data management is being done for all models at once, which is usually a good choice for me. Nevertheless, we need to remember 6 ways to start programs and how the different tasks depend on each other. This does not scale to serious projects!
To make it manageable, these templates
store source files in directories for each of the steps of the analysis,
store all outputs in a dedicated directory (tree),
provide a worked example using pytask to manage the workflow (with a directed acyclic graph in the background).
Directory Structure#
The big picture#
The following graph shows the contents of the example project root directory after
executing pytask, when you only select the Python example:
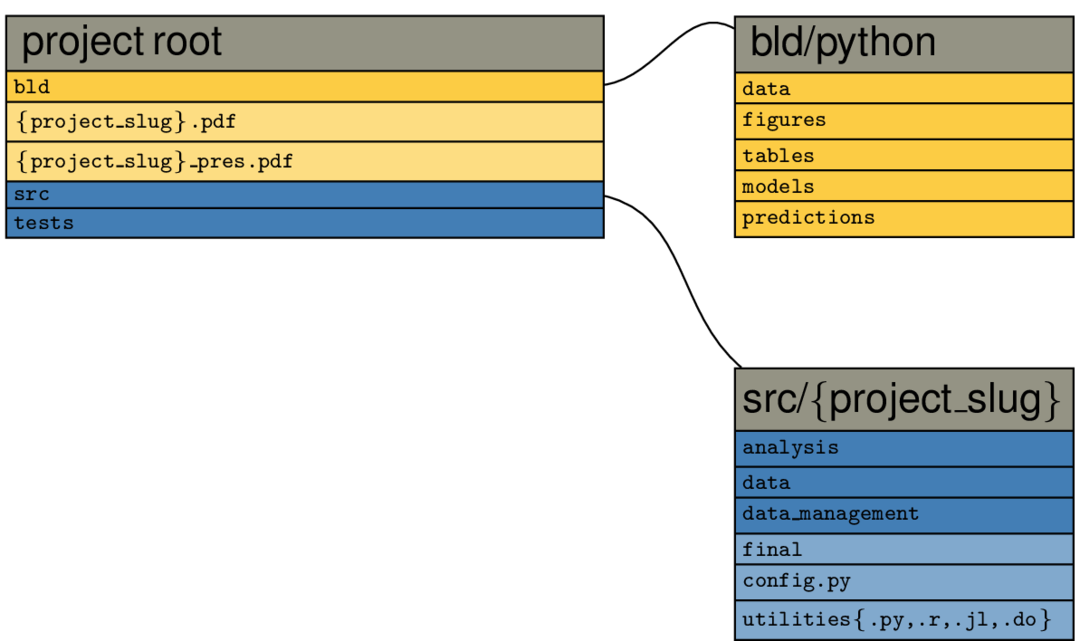
All other languages work accordingly, just replace python by the respective language
in the above path.
Files and directories in yellow are constructed by pytask; those with a bluish background are added directly by the researcher. You immediately see the separation of inputs and outputs (one of our guiding principles) at work:
All source code is in the src directory
All outputs are constructed in the bld directory
Note
The paper and presentation are copied to the root so they can be opened easily
The contents of both the root/bld and the root/src directories directly follow the steps of the analysis from the workflow section.
The idea is that everything that needs to be run during the, say, analysis step, is specified in root/src/analysis and all its output is placed in root/bld/analysis.
Some differences:
Because they are accessed frequently, figures and the like get extra directories in root/bld
The directory root/src contains many more subdirectories and files:
utilities.py provides code that may be used by different steps of the project. Little code snippets for input / output or stuff that is not directly related to the model would go here.
Zooming in#
Lets go one step deeper and consider the root/src directory in more detail:
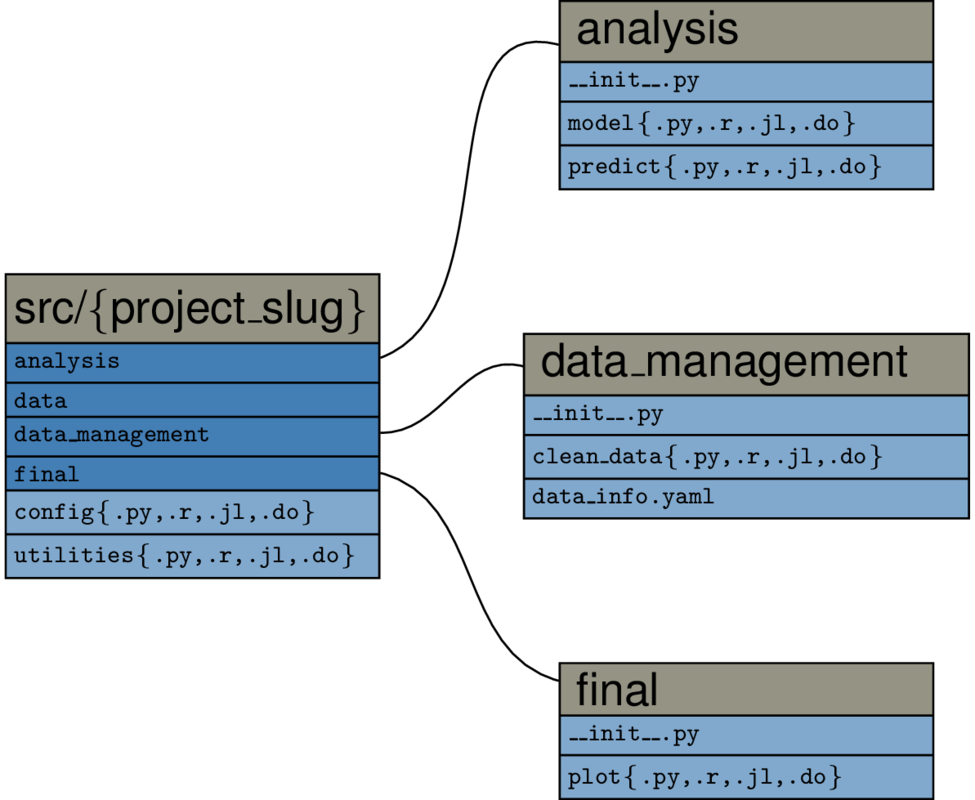
It is imperative that you do all the task handling inside the task_xxx.py-scripts,
using the pathlib library. This ensures that
your project can be used on different machines and it minimises the potential for
cross-platform errors.
For running Python source code from pytask, simply include depends_on and produces
as inputs to your function.
For running scripts in other languages, pass all required files (inputs, log files,
outputs) as arguments to the @pytask.mark.[x]-decorator. You can then read them in.
Check the sections on different programming languages for examples.
Directed Acyclic Graphs#
The way to specify dependencies between data, code and tasks to perform for a computer is a directed acyclic graph. A graph is simply a set of nodes (files, in our case) and edges that connect pairs of nodes (tasks to perform). Directed means that the order of how we connect a pair of nodes matters, we thus add arrows to all edges. Acyclic means that there are no directed cycles: When you traverse a graph in the direction of the arrows, there may not be a way to end up at the same node again.
This is the dependency graph of the example project (open the image in a different window to zoom in)
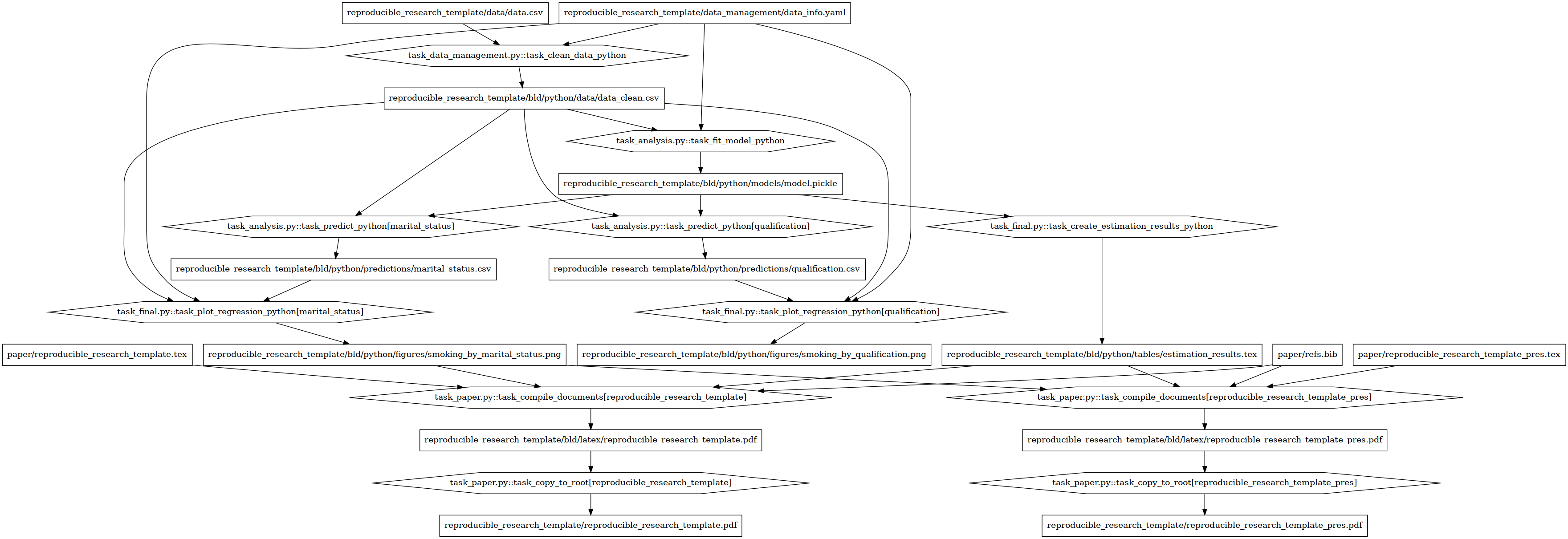
The nodes have different shapes in order to distinguish tasks from files. The rectangles denote targets or dependencies like figures, data sets or stored models. The hexagons denote task files. Even in this simple example project we already see that the dependency structure can be complex.
In a first run, all targets have to be generated, of course. In later runs, a target
only needs to be re-generated if one of its direct dependencies changes. E.g. when
we alter paper/research_pres_30min.tex (mid-right) we need to rebuild only the
presentation pdf file. If we alter rrt/data_management/data_info.yaml (top-right),
however, we need to rebuild everything. Note, that the only important thing at this
point is to understand the general idea.
Of course this is overkill for a simple example – we could easily keep the code closer together than this. But such a strategy does not scale to serious papers with many different specifications. As a case in point, consider the DAG for an early version of [vG15]:
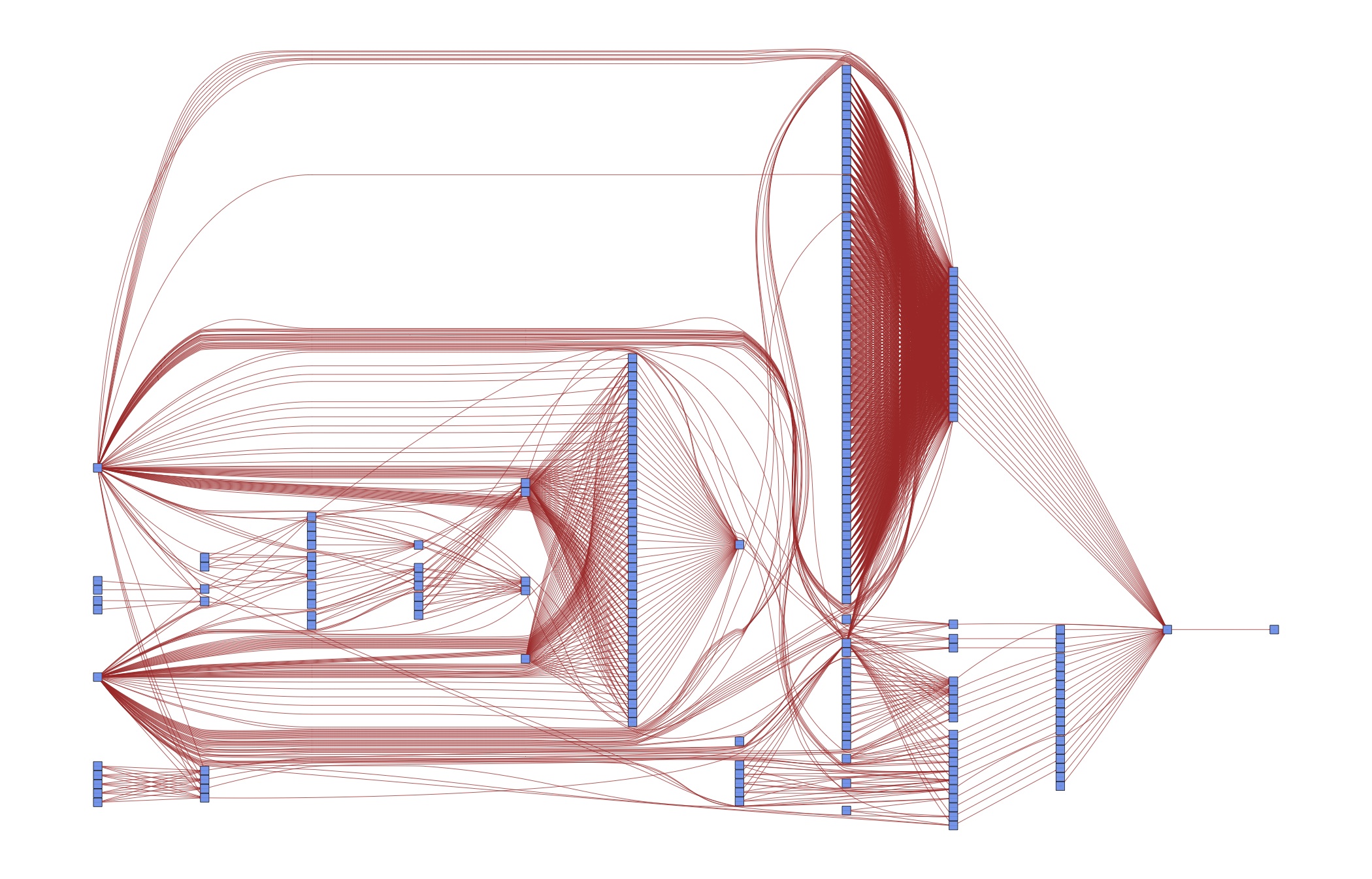
Do you want to keep those dependencies in your head? Or would it be useful to specify them once and for all in order to have more time for thinking about research? The next section shows you how to do that.
Introduction to pytask#
pytask is our tool of choice to automate the dependency tracking via a DAG (directed acyclic graph) structure. It has been written by Uni Bonn alumnus Tobias Raabe out of frustration with other tools.
pytask is inspired by pytest and leverages the same plugin system. If you are familiar with pytest, getting started with pytask should be a very smooth process.
pytask will look for Python scripts named task_[specifier].py in all subdirectories of
your project. Within those scripts, it will execute functions that start with task_.
Have a look at its excellent documentation. At present, there are additional plugins to run R scripts, Julia scripts, Stata do-files, and to compile documents via LaTeX.
We will have more to say about the directory structure in the Directory Structure
section. For now, we note that a step towards achieving the goal of clearly separating
inputs and outputs is that we specify a separate build directory. All output files go
there (including intermediate output), it is never kept under version control, and it
can be safely removed – everything in it will be reconstructed automatically the next
time you run pytask.
Pytask Overview#
From a high-level perspective, pytask works in the following way:
pytask reads your instructions and sets the build order.
Think of a dependency graph here.
pytask stops when it detects a circular dependency or ambiguous ways to build a target (e.g., you specify the same target twice).
Both are major advantages over a workflow script, let alone doing the dependency tracking in your mind.
pytask decides which tasks need to be executed and performs the required actions.
Minimal rebuilds are a huge speed gain compared to a workflow script.
These gains are large enough to make projects break or succeed.
We have just touched upon the tip of the iceberg here; pytask has many more goodies to offer. Its documentation is an excellent source.